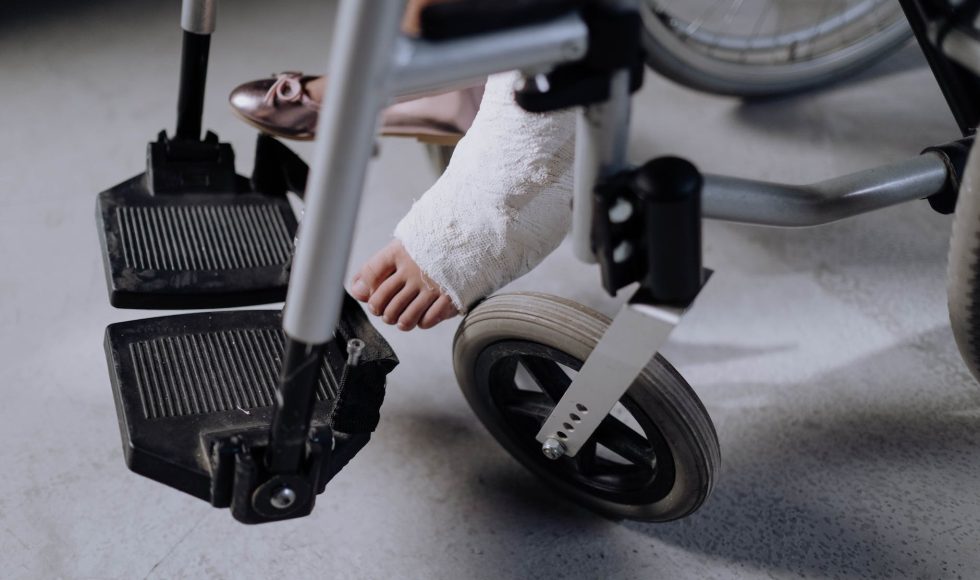
Teresa Street from the University of Oxford in the UK presented at the Nanopore Community Meeting 2019 on “Metagenomic nanopore sequencing for analysis of orthopedic device-related infection.” Street spoke about the impact of hip, knee, shoulder, elbow, and ankle replacements and the percentage of revisions due to the suspicion of infection. Further surgeries are often […]
Matthew Clark from The Genome Analysis Center presented at the Nanopore Community Meeting 2016 a session with the title “MinION metagenomics: from mock communities to clinical samples.” They took a mock community with twenty equimolar genomes. They were impressed with the results: 98%+ aligned, and there were long alignments. At the time, the group used […]
Justin O’Grady, Associate Professor in Medical Microbiology at the University of East Anglia, presented at the Nanopore Community Meeting 2017 on “Rapid metagenomic diagnosis of hospital acquired pneumonia.” They spoke about the “loss” of effective antibiotics because of rising resistance. O’Grady suggests using metagenomics-based infection diagnostics to use narrow-spectrum antibiotics. Nanopore sequencing is useful because […]
Brian Ondov from the National Biodefense Analysis and Countermeasures Centre (NBACC) spoke at the Nanopore Community Meeting 2015 on “Leveraging MinHash for rapid identification of nanopore data on mobile hardware.” Ondov explained how the portability of the MinION device can be paired with portable hardware. They noted that memory expansion on a small device can […]
Mads Albertsen, Associate Professor at Aalborg University, presented at the Nanopore Community Meeting 2017 on “Genome-centric metagenomics in the long-read era.” Albertsen talked about how we live in a bacterial world. However, only a small fraction can be grown. Single-cell genomics is challenging. Metagenomics has the potential to separate genomes through binning. Now you can […]
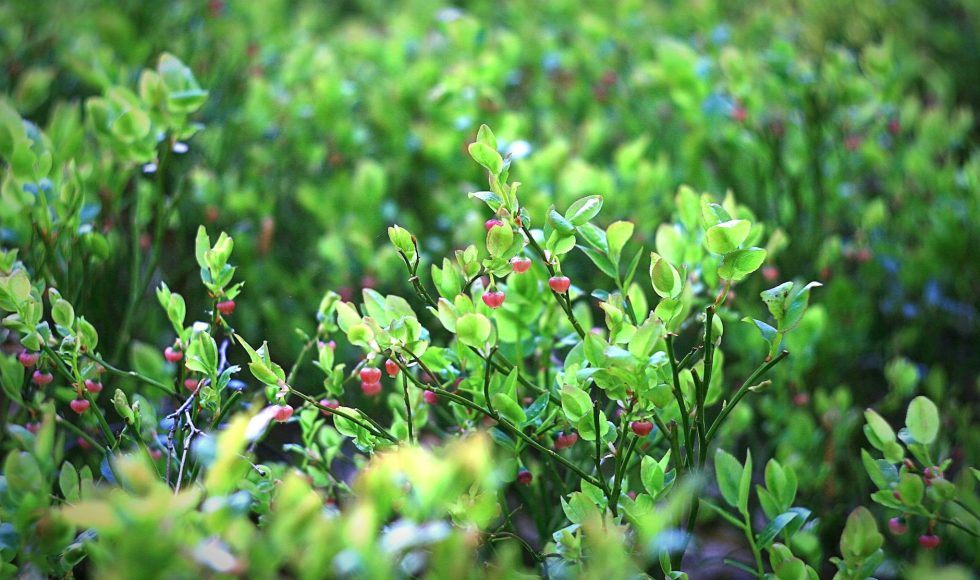
Seda Mirzoyan from Rutgers University presented at the Nanopore Community Meeting 2019 on “Using SIP and MinION sequencing to uncover active microbial communities in blueberry farm and forest soil systems.” Mirzoyan characterized active communities by using SIP and the MinION system. They spoke about the United States being the largest blueberry producer, with over fourteen […]
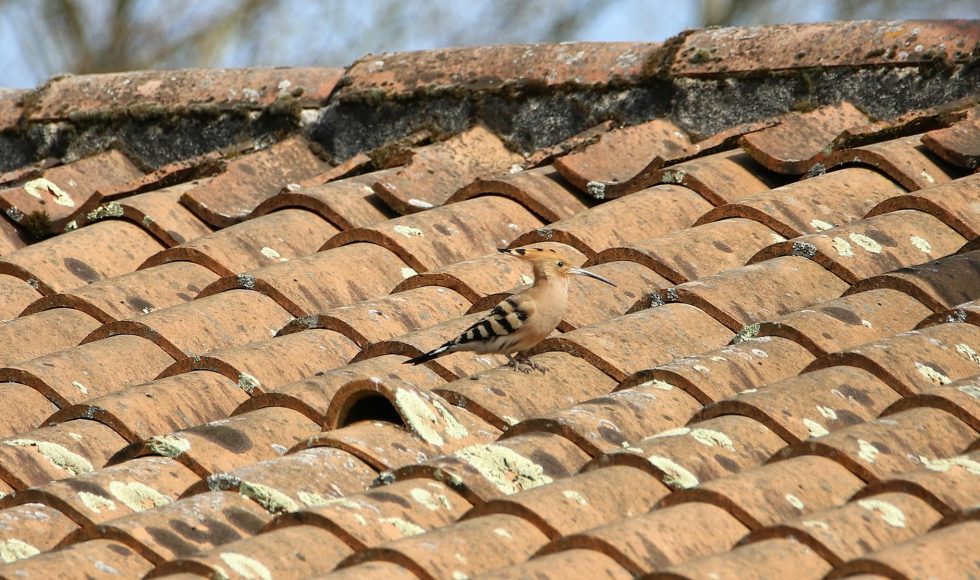
Lewis Stevens from Northwestern University presented at the Nanopore Community Meeting 2019 on “Reference genomes from the field: the genome of Caenorhabditis bovis.” Stevens spoke about nematodes and their importance as parasites! Nematodes are estimated to infect 1.5 billion people worldwide! Wow! C. elegans is, however, distantly related to the nematode parasites. C. bovis may […]
Ryan Wick from the University of Melbourne spoke at the Nanopore Community Meeting 2017. They explained that assemblies still fail. The title of the session is “Assembly is still hard: challenges in genome assembly in the era of long reads.” Wick shared an example of a bacterial genome sequence that leads to inaccuracy in sequencing. […]
Dylan Maghini from Stanford University presented at the Nanopore Community Meeting 2019 on “Genomes from metagenomes: assembling bacterial genomes with nanopore sequencing.” Maghini spoke about the importance of the human gut microbiome on human health. They noted that we still have an incomplete understanding of the gut microbiome. For example, de novo assembly aims to […]
Caitlin Singleton from Aalborg University in Denmark presented at the Nanopore Community Meeting 2020 a session on metagenomic analyses titled “From MAGs to riches: exploring the microbial community of activated sludge using 1,083 high-quality metagenome-assembled genomes.” Singleton is a postdoc and explained why activated sludge is important. Activated sludge is used to treat wastewater to […]