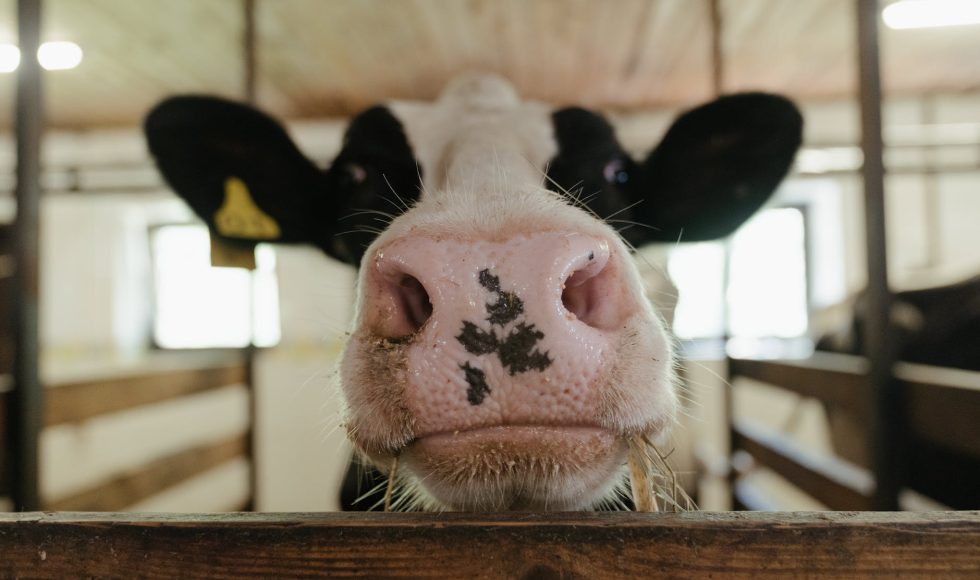
Tuan Viet Nguyen, a research scientist from Agriculture Victoria in Australia, presented at London Calling 2023 on “Discovering the missing variation: a long-read sequencing study into the structural variation in two dairy breeds.” They defined structural variants (SVs) as larger than 50 bps and can be deletions, duplications, and inversions. Structural variants, Nguyen noted, are […]
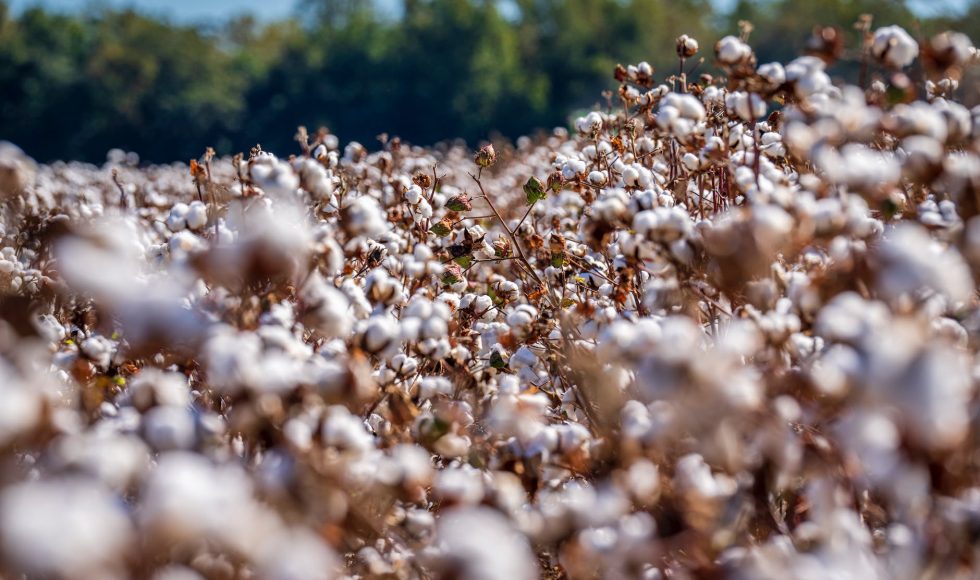
Babita Singh from the CSIR National Botanical Research Institute in India presented at London Calling 2023 on “Exploring targeted genetic diversity in the core Indian cotton germplasm using Oxford Nanopore platforms.” Singh is a postdoctoral fellow and spoke about how “cotton fibers are unicellular epidermal trichomes of ovules” and one of the most abundant crops […]
Tonight I watched the London Calling 2023 session entitled “Advancing targeted haplotyping in pharmacogenomics using adaptive sampling.” Koen Deserranno from Ghent University in Belgium spoke about their research. They are a pharmacist by training and are interested in pharmacogenomics (PGx): personalization of drug therapy based on genomic sequences of the individual. Deserranno spoke about how […]
Caroline Koch from the Imperial College London in the UK presented at London Calling 2023 about “Novel screening platform for highly multiplexed biomarker analysis.” Koch is a PH.D. student. They spoke about how humans express diseases differently: we have different genes and live different lives. Koch described the need for individual highly multiplexed platform to […]
Tonight I continued watching the London Calling 2023 Showcase Stage recordings for sessions on targeted sequencing. The speaker was Lukas Weilguny a Ph.D. student from the EMBL-EBI in the UK. The title of the session was “Dynamic, adaptive sampling during nanopore sequencing using Bayesian experimental design.” They worked on a model that takes the sequencing […]
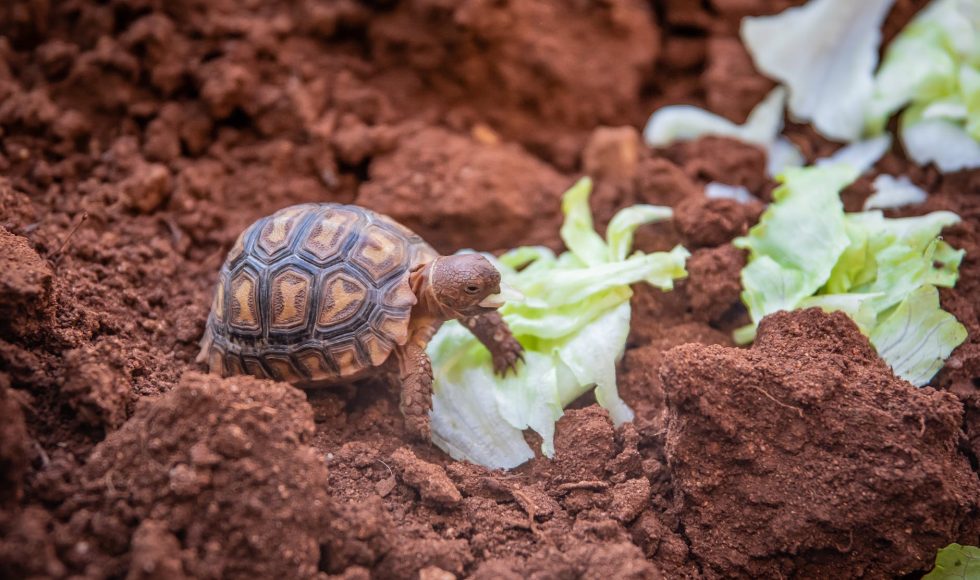
Tonight I watched the Oxford Nanopore Technologies (ONT) London Calling 2023 Showcase Stage on Conservation. In this session, Luca Pandolfini from the Italian Institute of Technology in Italy presented “From genome assembly to epigenome characterisation: a nanopore journey in the footsteps of an endangered tortoise.” Pyxis arachnoides is a tiny tortoise that is critically endangered […]
Cecilia C.S. Yeung & Olga Sala-Torra from the Fred Hutchinson Cancer Center presented the session “Potential clinical utility of long-read sequencing in myeloid neoplasms.” They are both clinicians and saw the potential of long-read sequencing for detection of myeloid neoplasms. The assay has the potential to rapid detect cytogenetic aberrations. Sala-Torra spoke about how fusions […]
Tonight I continued watching the London Calling 2023 targeted sequencing showcase. In the four-minute recording entitled “The potential use of nanopore sequencing for molecular diagnosis of germline cancer predisposition,” Romain Boidot from the Centre Georges-François Leclerc in France spoke about their use of Oxford Nanopore Technologies (ONT) for molecular diagnosis and sequencing. Their current analysis […]
Tonight I watched the London Calling 2023 session on targeted sequencing. The first question asked was why select targeted sequencing. One panelist spoke about the ability to try the technology and modify it for their needs. Another speaker explained that creating a custom panel would be easier for the analysis. A third panelist said that […]
Janessa Laskin from Canada’s Michael Smith Genome Sciences Centre at BC Cancer presented at London Calling 2023 on the “Potential use of nanopore sequencing in clinical cancer genomics.” Laskin is a clinician and medical oncologist. They spoke about the uncertainty of what treatments to select for patients and the idea of personalized medicine. Laskin explained […]