Noah Bryan from the Bayview Secondary School in Canada presented at the Nanopore Community Meeting in Boston. The title of the five-minute session was “Is the water safe to drink/ The rapid test is the missing link!” Bryan is a sixteen-year-old student grade twelve student from Toronto, Canada. They developed a water test using the […]
I continued watching the “Enhanced microbial profiling and metagenomics with nanopore sequencing” webinar. The second speaker was Steven Batinovic, a Field Application Scientist with Oxford Nanopore Technologies (ONT). The title of their session was “What you’re missing matters: delivering the future of microbial genomics with Oxford Nanopore.” Batinovic started by listing the microbiology and infectious […]
Tonight, I watched a Nanopore webinar by Tianyuan Zhang. The session’s title is “The Newest Oxford Nanopore R10.4.1 Full-length 16S rRNA Sequencing Enables the Accurate Resolution of Species-Level Microbial Community Profiling.” Zhang spoke about the resolution gains with full-length 16S sequencing, citing a Nature article from 2019. However, the error rate of ONT was previously […]
Trish Simner is the Director of Bacteriology and Infectious Disease Sequencing Laboratories. They work at the Johns Hopkins University School of Medicine in Baltimore, Maryland. They presented at London Calling 2024. The session’s title was “Bringing Nanopore Sequencing into the Clinical Microbiology Setting with Targeted Approaches.” Simner spoke about the advantages of long reads and […]
Tonight, I watched the London Calling 2024 EPI2ME Product Demo. The session title is: “EPI2ME: democratising bioinformatics – from point-and-click analysis to custom integrations.” Sarah Griffiths, ONT Bioinformatics Workflow Developer, gave an overview of the EPI2ME workflows. The workflows use Nexflow and containers. At the time of recording, they had seventeen workflows. Griffiths noted that […]
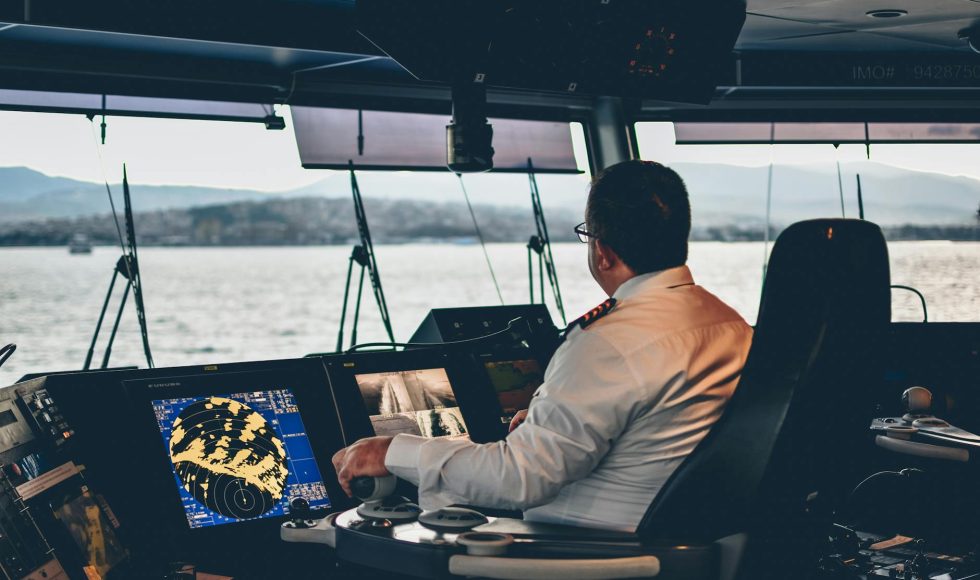
Sophie Colston from the US Naval Research Laboratory presented at London Calling 2019 on “Field forward sequencing in naval environments.” The title of this session caught my attention as it aligns well with the course I am developing. Colston is a microbiologist. The Naval Research Lab (NRL) supports research and technological development. The NRL has […]
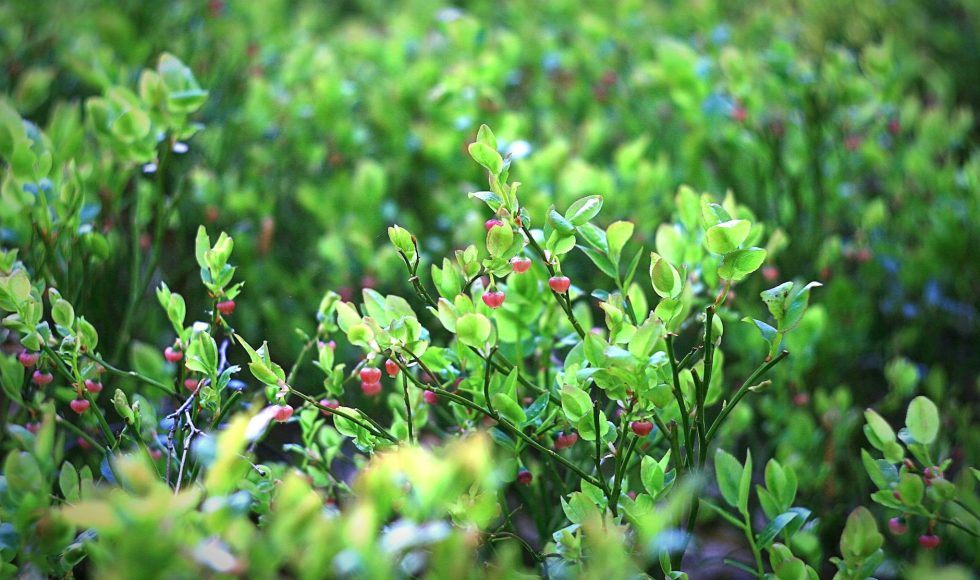
Seda Mirzoyan from Rutgers University presented at the Nanopore Community Meeting 2019 on “Using SIP and MinION sequencing to uncover active microbial communities in blueberry farm and forest soil systems.” Mirzoyan characterized active communities by using SIP and the MinION system. They spoke about the United States being the largest blueberry producer, with over fourteen […]
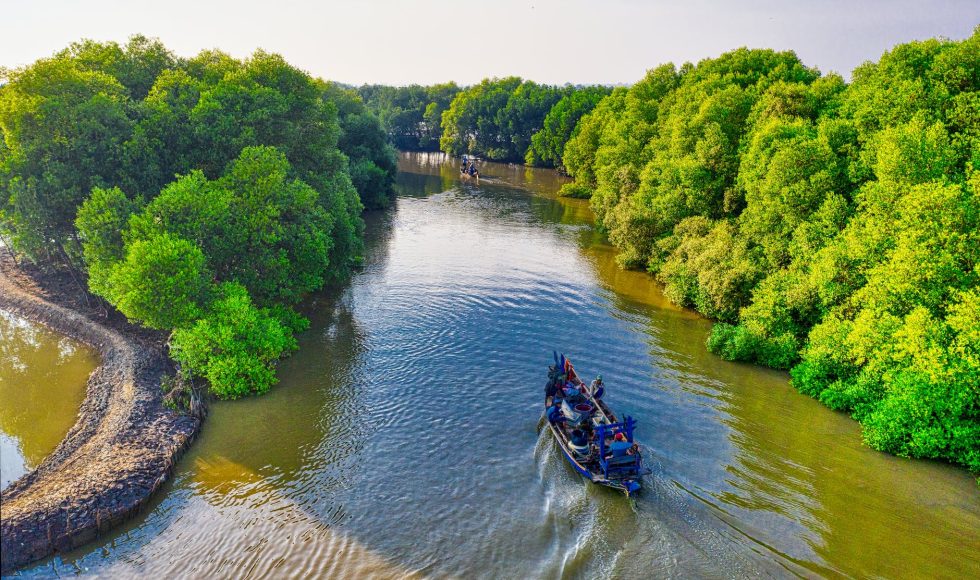
Felipe Baez Aguirre from the University of the Andes in Colombia presented a session entitled “On-site nanopore sequencing reveals microbial diversity of a Colombian Pacific Coast mangrove soils” at the Nanopore Community Meeting in Houston. Baez Aguirre spoke about the location of the studies. They had limited access to electricity in the Bahia Malaga. The […]
Tonight I watched the Update from Oxford Nanopore Technologies from NCM Houston. They started by explaining how the engineered motor protein and nanopore have been optimized for single-molecule high-throughput sequencing. There is an RNA-specific motor protein and nanopore. ONT allows for RNA modification detection, isoform characterization, and comparison of tails. RNA 004 is the new […]
Tim Walker from the Technical Services Team with Oxford Nanopore Technologies presented the Nanopore Learning Course – Metagenomics lesson I watched tonight. They spoke about metagenomic classification techniques to identify organisms from sequence data and annotate genes. For targeted metagenomics, ITS, 16S, or antibiotic resistance genes can be amplified and sequenced. Whole genome sequencing can […]