Seda Mirzoyan from Rutgers University presented at the Nanopore Community Meeting 2019 on “Using SIP and MinION sequencing to uncover active microbial communities in blueberry farm and forest soil systems.” Mirzoyan characterized active communities by using SIP and the MinION system. They spoke about the United States being the largest blueberry producer, with over fourteen […]
Lewis Stevens from Northwestern University presented at the Nanopore Community Meeting 2019 on “Reference genomes from the field: the genome of Caenorhabditis bovis.” Stevens spoke about nematodes and their importance as parasites! Nematodes are estimated to infect 1.5 billion people worldwide! Wow! C. elegans is, however, distantly related to the nematode parasites. C. bovis may […]
Ryan Wick from the University of Melbourne spoke at the Nanopore Community Meeting 2017. They explained that assemblies still fail. The title of the session is “Assembly is still hard: challenges in genome assembly in the era of long reads.” Wick shared an example of a bacterial genome sequence that leads to inaccuracy in sequencing. […]
Dylan Maghini from Stanford University presented at the Nanopore Community Meeting 2019 on “Genomes from metagenomes: assembling bacterial genomes with nanopore sequencing.” Maghini spoke about the importance of the human gut microbiome on human health. They noted that we still have an incomplete understanding of the gut microbiome. For example, de novo assembly aims to […]
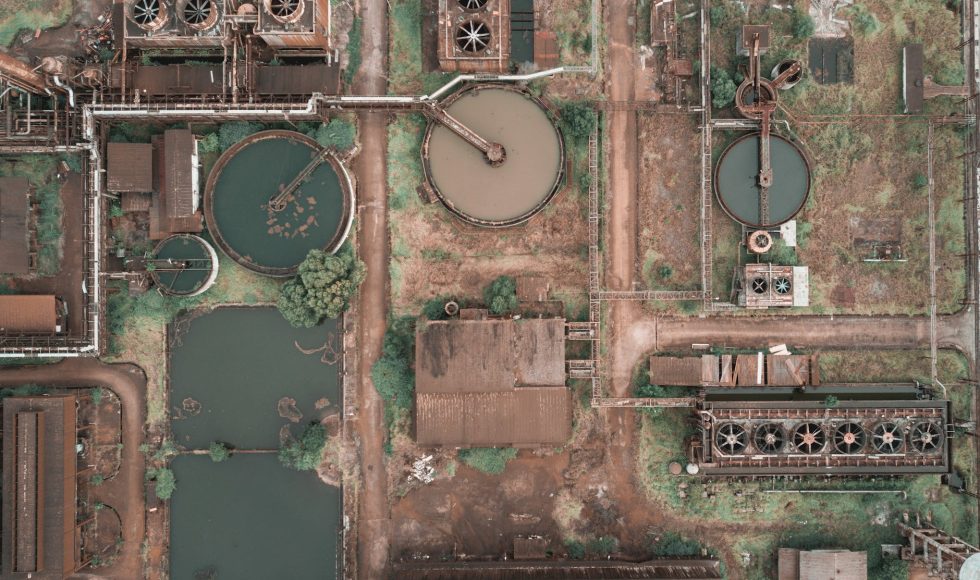
Caitlin Singleton from Aalborg University in Denmark presented at the Nanopore Community Meeting 2020 a session on metagenomic analyses titled “From MAGs to riches: exploring the microbial community of activated sludge using 1,083 high-quality metagenome-assembled genomes.” Singleton is a postdoc and explained why activated sludge is important. Activated sludge is used to treat wastewater to […]
Mantas Sereika from Aalborg University in Germany presented at the Nanopore Community Meeting 2021 a session with a title that caught my attention tonight: “Nanopore R10.4 enables near-perfect bacterial genomes.” They spoke about Nanopore sequencing raw read accuracy improvements and issues with homopolymers. Insertions and deletions in homopolymer regions can be an issue causing frameshifts. […]
This afternoon, I came back from visiting KBase in Berkeley. I am still watching sessions from previous London Callings to learn about different topics I will discuss in the upcoming Portable Genome Sequencing course. Tonight, I found the session by David R. Greig from Public Health England in the UK. Their 2021 London Calling session […]
I am starting to watch older Oxford Nanopore Technologies (ONT) videos to prepare background information for the course we are developing. Tonight, I watched a session by Rob Harbert from Stonehill College. The title of the presentation was “Monitoring plant biodiversity in aquatic eDNA with low-cost Oxford Nanopore Flongle sequencing.”They used the Flongle for metabarcoding. […]
Today, I spent time with the KBase Ed group at UC Berkeley! The first day was full of tips and opportunities. As I prepare for the Portable Genome Sequencing course later this semester, I am trying to learn about using Oxford Nanopore Technologies (ONT) devices for sequencing plasmids, microbes, metagenomes, and transcriptomes. Tonight, I watched […]
Tonight, I was searching for videos for the course I am preparing to teach on portable genome sequencing technologies. I want to discuss the use of long-read sequencing for identifying and characterizing native plasmids. I found an older yet very relevant London Calling 2019 session. Severine Rangama from the University of Warwick presented a short […]