Brian Ondov from the National Biodefense Analysis and Countermeasures Centre (NBACC) spoke at the Nanopore Community Meeting 2015 on “Leveraging MinHash for rapid identification of nanopore data on mobile hardware.” Ondov explained how the portability of the MinION device can be paired with portable hardware. They noted that memory expansion on a small device can […]
Mads Albertsen, Associate Professor at Aalborg University, presented at the Nanopore Community Meeting 2017 on “Genome-centric metagenomics in the long-read era.” Albertsen talked about how we live in a bacterial world. However, only a small fraction can be grown. Single-cell genomics is challenging. Metagenomics has the potential to separate genomes through binning. Now you can […]
Seda Mirzoyan from Rutgers University presented at the Nanopore Community Meeting 2019 on “Using SIP and MinION sequencing to uncover active microbial communities in blueberry farm and forest soil systems.” Mirzoyan characterized active communities by using SIP and the MinION system. They spoke about the United States being the largest blueberry producer, with over fourteen […]
Ryan Wick from the University of Melbourne spoke at the Nanopore Community Meeting 2017. They explained that assemblies still fail. The title of the session is “Assembly is still hard: challenges in genome assembly in the era of long reads.” Wick shared an example of a bacterial genome sequence that leads to inaccuracy in sequencing. […]
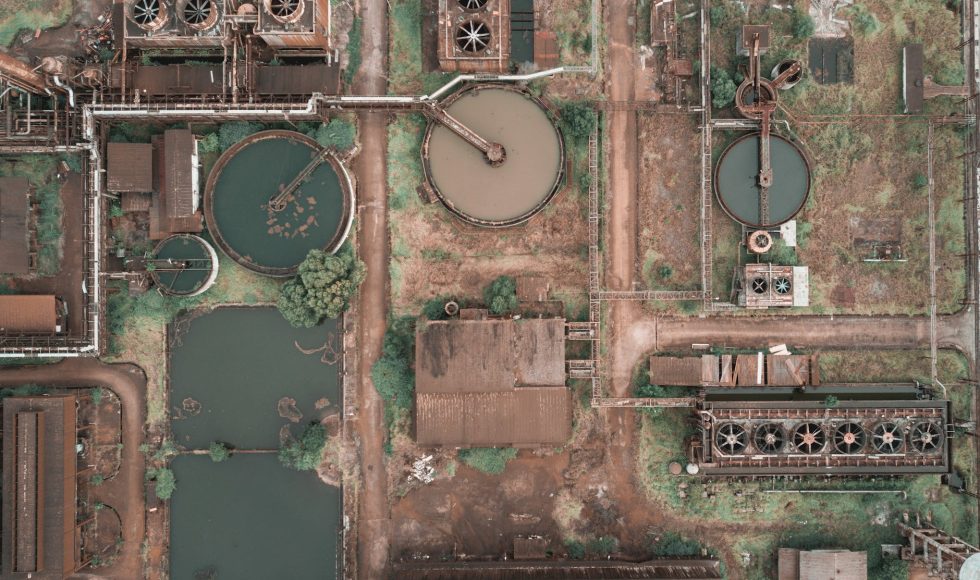
Caitlin Singleton from Aalborg University in Denmark presented at the Nanopore Community Meeting 2020 a session on metagenomic analyses titled “From MAGs to riches: exploring the microbial community of activated sludge using 1,083 high-quality metagenome-assembled genomes.” Singleton is a postdoc and explained why activated sludge is important. Activated sludge is used to treat wastewater to […]
This afternoon, I came back from visiting KBase in Berkeley. I am still watching sessions from previous London Callings to learn about different topics I will discuss in the upcoming Portable Genome Sequencing course. Tonight, I found the session by David R. Greig from Public Health England in the UK. Their 2021 London Calling session […]
I am starting to watch older Oxford Nanopore Technologies (ONT) videos to prepare background information for the course we are developing. Tonight, I watched a session by Rob Harbert from Stonehill College. The title of the presentation was “Monitoring plant biodiversity in aquatic eDNA with low-cost Oxford Nanopore Flongle sequencing.”They used the Flongle for metabarcoding. […]
Today, I spent time with the KBase Ed group at UC Berkeley! The first day was full of tips and opportunities. As I prepare for the Portable Genome Sequencing course later this semester, I am trying to learn about using Oxford Nanopore Technologies (ONT) devices for sequencing plasmids, microbes, metagenomes, and transcriptomes. Tonight, I watched […]
Tonight, I was searching for videos for the course I am preparing to teach on portable genome sequencing technologies. I want to discuss the use of long-read sequencing for identifying and characterizing native plasmids. I found an older yet very relevant London Calling 2019 session. Severine Rangama from the University of Warwick presented a short […]
Colette Felton from the University of California at Santa Cruz presented at the Nanopore Community Meeting in Houston about “Haplotypes, isoforms, and fusions: towards a richer cancer transcriptome.” The Felton read studies splicing, and long reads could be used to detect isoforms. The lab developed a tool called FLAIR2 to study differential splicing. Felton’s lab […]