Mike Clark from the University of Melbourne in Australia presented at London Calling 2019 on “Deep transcriptomic sampling with long-read single cell RNA sequencing.” Clark gave the first talk in the session and explained the power of expression profiles of single cells (scRNA-seq) to identify cell types and variations in gene expression. scRNA-seq can be […]
My Linh Thibodeau from the University of British Columbia in Canada presented at London Calling 2019 on “Resolution of germline hereditary cancer structural variants using nanopore sequencing.” They began talking about the Personalized OncoGenomics Program (POG), which is an initiative of BC Cancer. The study enrolls participants and conducts extensive genomic analyses. The team evaluated […]
Pore-C is a method I hear about, yet I don’t fully understand the details. Eoghan Harrington from Oxford Nanopore Technologies (ONT) presented at London Calling 2019 a session titled “Pore-C: a method for genome-wide, multi-contact chromosome conformation capture.”Harrington is part of the applications team and focuses on genomic applications. They collaborate with various partners. The […]
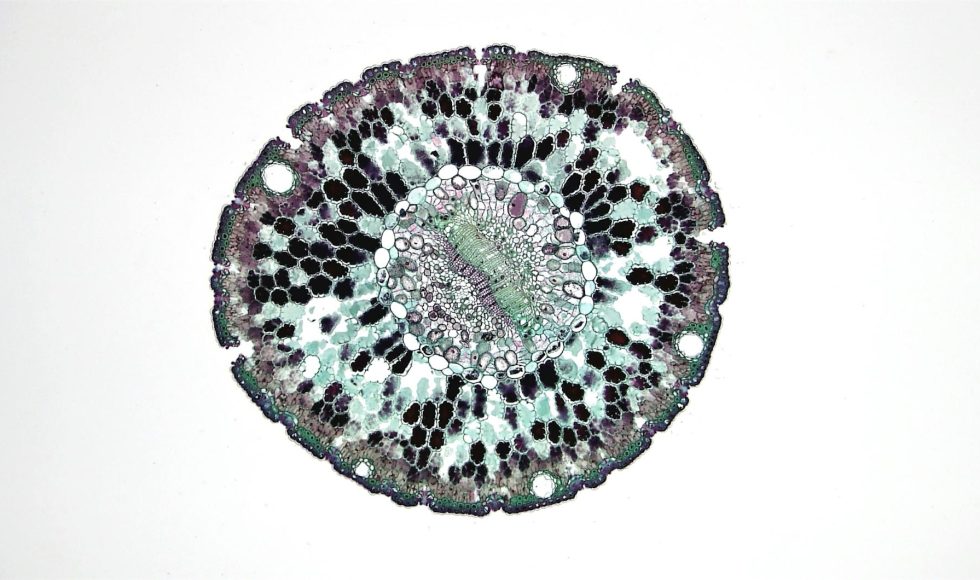
Marie-Christine Carpentier, from the Genome and Plant Development Laboratory in France, presented at London Calling 2019 on “Using long-read nanopore sequencing to unravel structural variations in plants.” We have been trying plant genome sequencing and want to learn more. Carpentier spoke about transposable elements. Mobile elements can move within a genome. Carpentier noted that the […]
Yusmiati Liau from the University of Otago in New Zealand spoke at London Calling 2019 about “Nanopore sequencing of the CYP2D6 pharmacogene.” Liau explained that pharmacogenetics is used to learn about genes that affect responses to drugs. There is a standard nomenclature for pharmacogenetics. The PHarmVar database was used as an example. The CY2PD6 gene […]
David Greig from Public Health England presented a session in London Calling 2019 with the title “Comparison of single nucleotide variants identified by Illumina and Oxford Nanopore technologies in the context of a potential outbreak of Shiga toxin producing.” They explained how they work on pathogen surveillance and focus on STEC: Shiga toxin-producing Escherichia coli. […]
Monica Kehoe from the Department of Primary Industries and Regional Development in Australia presented at the 2019 London Calling meeting. The session’s title was “Nanopore sequencing and analysis of plant pathogenic viruses—more than just rapid diagnostics?” Kehoe is based in Western Australia and using Nanopore sequencing to sequence RNA plant viruses. Their focus is to […]
Mark T.W. Ebbert from the Mayo Clinic presented at London Calling 2019 on “Long-read sequencing technologies resolve most ‘dark’ and ‘camouflaged’ gene regions.”Dark and camouflaged regions? Ebbert explained that regions can be dark because there are no reads available (“dark by depth“) or dark by low sequence quality (“dark by MAPQ“). Ebbert explained that most […]
Stella Loke from Deakin University in Australia spoke at London Calling 2019 about “Optimising plant DNA extraction for nanopore sequencing.” We are considering sequencing plant DNA this summer and, therefore, want to learn more about plant DNA challenges. Loke spoke about wanting the nano spec readings from Qubit to match the nano spectrophotometer. Loke confirms […]
Amanda Warr from The Roslin Institute in the UK presented at London Calling 2019, “Going full circle: assembly of high-quality, single-contig microbial genomes from the rumen microbiome using long-read sequencing.” I had not watched this session before! They spoke about the rumen: a specialized stomach with a complex microbial environment. They are interested in the […]