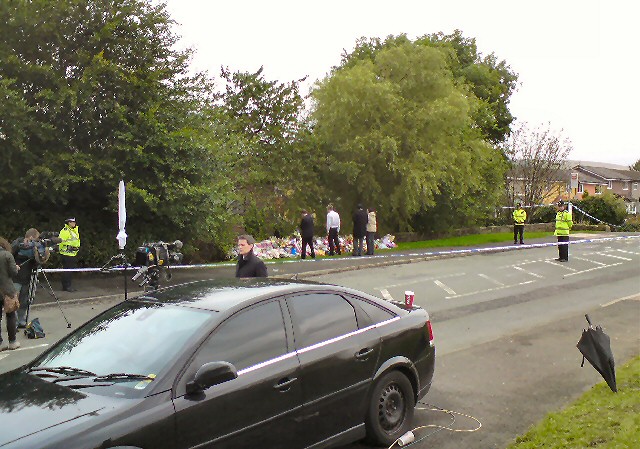
Abderaouf Hamza from the Institut Curie in France spoke at the Nanopore London Calling event last year about “Targeted nanopore sequencing ushers in the era of routine long-read sequencing in translational research laboratories.” This ten-minute session started with how Hamza uses nanopore sequencing for cancer genomics, particularly using adaptive sampling. They use target region enrichment […]
Tonight I watched a five-minute Oxford Nanopore Technologies London Calling 2022 session by Mrinalini Erkenswick Watsa and titled “Genomics in the jungle: a field laboratory success story.” Mrinalini Erkenswick Watsa is at the San Diego Zoo Wildlife Alliance and is a field scientist working on conservation. They spoke about the need to assess populations pre […]
Nicola Hall from the University of Oxford in the UK spoke at London Calling 2022 about “Multiplexed, long-read CaptureSeq identifies full-length transcript isoforms in the human brain.” I have been interested in CaptureSeq and this ten-minute session was intriguing. Hall and colleagues are interested in voltage-gated channels, but they have very low levels of RNA […]
Belen de la Morena Barrio from the University of Murcia/CIBERER, Spain presented at London Calling 2022 on “Genetic dissection of structural variants in study subjects with antithrombin deficiency by nanopore sequencing.” Morena Barrio was introduced to Nanopore during their doctoral work, and they have now developed methods. They spoke about thrombin deficiency: antithrombin deficiency (SERPINC1) […]
I finished the Nanopore Community Meeting 2022 videos and started the 2022 London Calling series. Bryant Catano is a Technical Applications Specialist with Oxford Nanopore Technologies. Catano is based at the New York offices. Their session was a masterclass on “Choosing the right nanopore sequencing device for your needs.” Catano spoke about the throughput of […]
Courtney Hall from the University of North Texas Health Science Center presented at the Nanopore Community Meeting 2022 on “STRspy-ing hidden variation in forensic DNA profiles using the MinION device.” Hall spoke about how short tendem repeats (STRs) are often used as the “gold standard for human identification in forensic investigations” due to their variability […]
Kame A. Galan-Huerta from the Autonomous University of Nuevo Leon in Mexico presented at the Nanopore Community Meeting 2022 a ten minute session entitled “Identification of resistance mutations to direct-acting antiviral agents against HCV in infected subjects in Mexico.” They spoke about adapting protocols to detect the RNA positive sense hepatitis C virus. Glan-Huerta explained […]
Tonight I watched Fritz Sedlazeck from The Baylor College of Medicine Human Genome Sequencing Center present at the Nanopore Community Meeting 2022 on “Rapid structural variant calling across AllOfUs using Oxford Nanopore sequencing.” Sedlazeck spoke about structural variations (SV) that are 50bp+ genomic alterations. Long read sequencing SV calling improves detection. Sedlazeck and team created […]
Melanie Sagniez from the CHU Sainte-Justine Research Centre in Canada presented on “Rapid leukemia classification using nanopore sequencing” as part of the Nanopore Community Meeting 2022. This ten-minute session described part of their Ph.D. project research. They first spoke about pediatric leukemia and the current diagnostic process. I didn’t realize that the process takes on […]
Mantas Sereika from Aalborg University in Denmark presented at the nanopore Community Meeting 2022 on “Targeted deep metagenomics for the recovery of novel closed microbial genomes from highly complex communities.” I thought this ten-minute session had an intriguing title, as I begin to wonder what we will do next in the BIT 295 course in […]